Genetic Investigations of SARS-CoV-2
The public health response to the SARS-CoV-2 virus is continually challenged by the evolution of new variants. As variants emerge, they are assessed by the World Health Organization to determine if they should be designated as a variant of concern (VOC). The potential of novel VOCs to cause new outbreaks highlights a heightened need for new antiviral therapeutic targets.
Small, noncoding RNAs of the host, including transfer RNA-derived RNAs (tDRs), have been recently identified as regulators of viral pathogenesis. These researchers have previously reported on a tDR regulatory binding site in SARS-CoV-2 that is mutated in several VOCs. Host genes are also targeted by tDRs, however, host genes targeted by tDRs in SARS-CoV-2 infected cells have not been identified at this time. New understanding of the role of tDRs in regulating the host response to viral infection will provide needed information for the future development of novel therapeutics.
MSI PI Lynne Bemis (professor, Biomedical Sciences, U of M Medical School Duluth Campus), Dr. Marna Ericson (The Hormel Institute), and Dr. Thomas Kono (Bioinformatics, MSI) are working on a project called “Sponging up tRNA derived RNAs (tDRs) implicated in the regulation of SARS-CoV-2.” This research team is working to gain understanding of the host genes regulated by two tDRs (tDR-Gly and tDR-Val), which also bind sequences within the SARS-CoV-2 genome. The project uses RNA-seq analysis to identify host genes with altered expression when the tDRs are depleted.
This project recently received a Research Computing Seed Grant. Professor Bemis uses MSI resources for research into identifying extracellular RNAs; the group is analyzing deep sequencing data that is already available and then developing new deep sequencing libraries from cell culture models. The Bemis group is also looking at the extracellular RNAs found in co-cultures of cancer cells and bacterial cells.
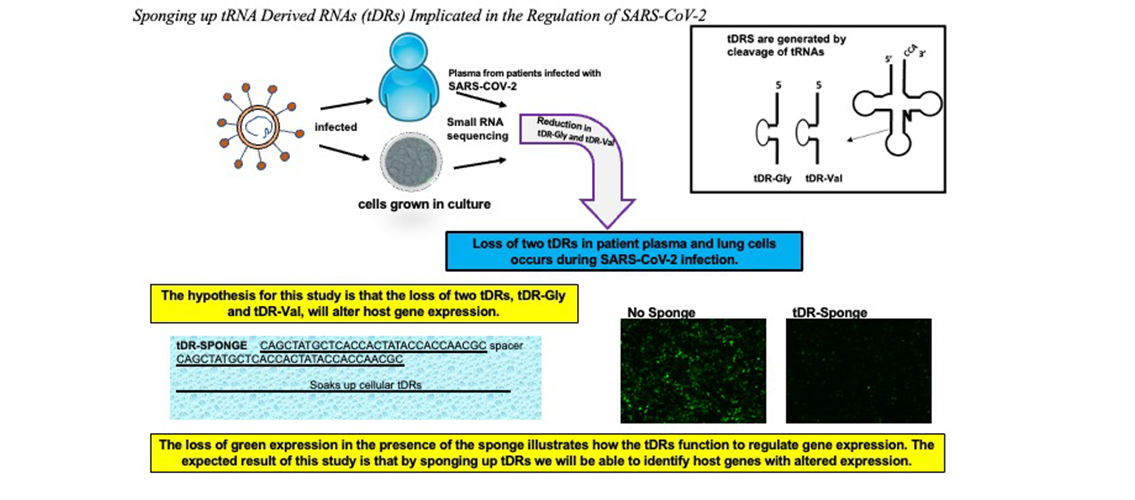
Image description: This project is based on previous work by the Bemis lab showing that two tDRs are downregulated following infection of cells by SARS-CoV-2. They also showed that sponge constructs are able to soak up/bind the host cellular tDRs and suppress gene expression. RNA was extracted from human cells transfected with sponge constructs and sequenced with high throughput sequencing technology. This project will analyze the RNA-seq data to identify host gene expression that changes in the presence of the tDR sponges.